The fox, the crow and the superbug: how wildlife could be an ‘early warning system’ for antibiotic resistance
A new study suggests wildlife can harbour drug-resistant bacteria, forming a feedback loop driving hard-to-treat infections in people.
- 29 April 2026
- 5 min read
- by Priya Joi

At a glance
- Researchers found a high-risk clone of Klebsiella pneumoniae, a bacterium that causes severe infections in hospitalised patients, in foxes and waterbirds across northern Italy.
- Every K. pneumoniae isolate from wildlife was resistant to antibiotics used to treat pneumonia, sepsis and meningitis, compared to roughly one in five clinical isolates from human patients.
- The team says monitoring wildlife could function as an early warning system, flagging the spread of resistance into ecosystems before it reaches clinical settings.
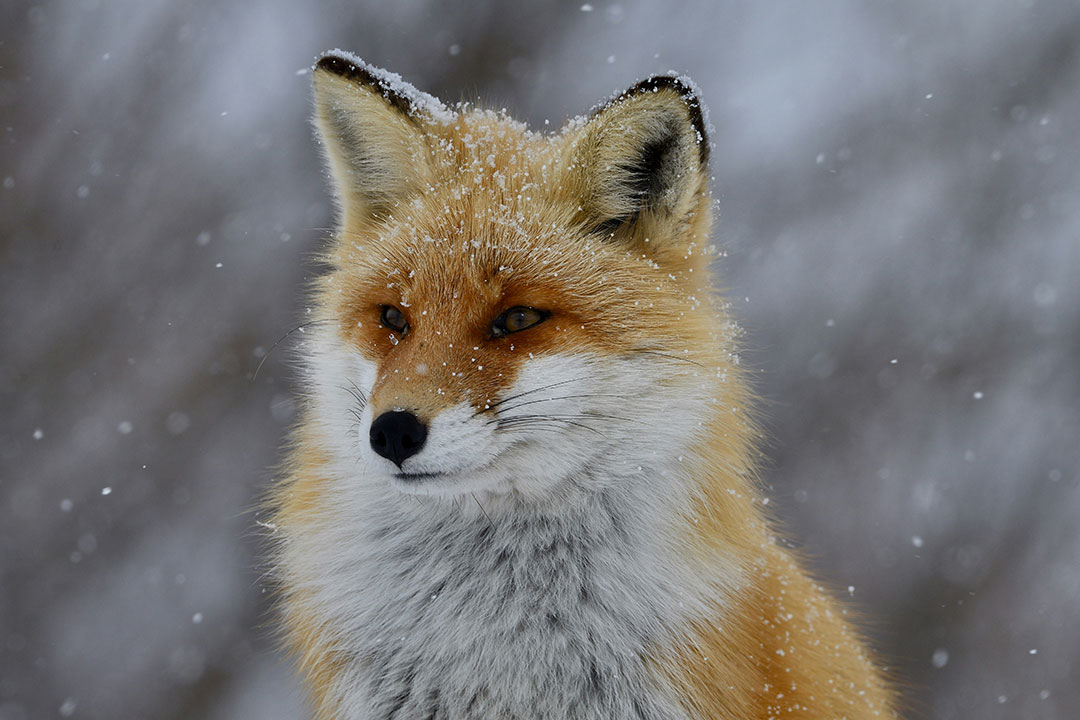
A red fox trotting through farmland on the outskirts of Parma may not seem the obvious place to look for bacteria that resist the most powerful antibiotics in a hospital formulary. But Italian researchers suggest that wildlife is exactly where we should be looking.
They’ve discovered antibiotic-resistant Klebsiella pneumoniae bacteria, which can cause urinary tract, respiratory tract and bloodstream infections, in foxes, magpies and crows at five times the levels found in hospitals.
One bacterial isolate was resistant to a ‘last-resort’ antibiotic that doctors use when nothing else works.
Lead author Prof Mauro Conter, at the University of Parma, told VaccinesWork that their research, published in Frontiers in Microbiology, indicates “potential spread from human or clinical sources into the environment”.
Low prevalence but high resistance levels, up to six antibiotic classes, underscores wildlife’s role as an under-monitored AMR reservoir.
This can create a concerning feedback loop, since wild animals act as reservoirs transferring resistant bacteria back to domestic animals and people.
In response, Conter and his team are calling for wildlife monitoring of antimicrobial resistance (AMR) to be built into national One Health strategies, and to track the transmission of resistance in a way that integrates wildlife, environmental and clinical sampling.
Why look for clinical bacteria in wild animals?
Antimicrobial resistance can be challenging to fight because resistance genes in microbes can hop between species on mobile genetic elements such as plasmids.
That means a resistant strain circulating in wastewater, agricultural run-off or urban waste can be picked up by animals foraging in these environments, even if those animals never come near a hospital or clinic.
K pneumoniae is one of the ESKAPE pathogens, a group of bacteria notorious for evading antibiotics and causing hard-to-treat infections in hospitals. This bacterium had already been found widely in wastewater, saltwater and freshwater environments, as well as in fresh vegetable production sites.
Have you read?
The researchers decided to examine whether wild animals could also play a central role in the spread of resistance genes. Foxes are well suited for this spread because they scavenge across urban, rural and wild landscapes, picking up bacteria from multiple sources and depositing them across short distances. Birds can do the same thing over hundreds of kilometres.
What did the researchers find?
The team at the University of Parma and the Istituto Zooprofilattico Sperimentale in Bologna collected faecal samples from 493 wild animals across the Emilia-Romagna region of northern Italy: 184 red foxes, 209 crows and magpies and 100 waterbirds including herons, swans and flamingos.
All had died from trauma or predation and, as they were wild animals, none would ever have been treated with antibiotics, indicating that the resistant bacteria originated elsewhere.
The researchers found Klebsiella species bacteria in 32 of 493 samples, and K. pneumoniae specifically in 10 samples, mainly in waterbirds and foxes.
Whole genome sequencing confirmed that all ten K. pneumoniae isolates belonged to ST307, a clone already well known in hospital outbreaks worldwide.
Every K. pneumoniae isolate was resistant to cephalosporins, a class of antibiotics used to treat pneumonia, sepsis and meningitis.
According to the latest European Centre for Disease Prevention and Control surveillance data, just 19.6% of K. pneumoniae isolates from human patients in Italy showed the same resistance. At the EU level, it was 9.3%.
Worryingly, one isolate from a fox also carried the NDM-5 carbapenemase gene, which can neutralise carbapenems, the antibiotics doctors turn to when nothing else works.
The pattern also held for fluoroquinolones, antibiotics used to treat severe urinary tract infections and pneumonia: 100% resistance in wildlife against 17.4% in Italian clinical isolates. Resistance to gentamicin, an aminoglycoside, reached 90% in the wildlife samples compared to 11.6% in human patients.
“The detection of the high-risk K. pneumoniae ST307 clone in wildlife samples was particularly concerning,” said Conter.
“This clone, carrying ESBL and carbapenemase genes on mobile plasmids, suggests potential spread from human and clinical sources into the environment via foxes and birds, which act as short- and long-distance disseminators.”
How are these resistant bacteria reaching wildlife?
The researchers believe the most likely route is environmental contamination. Wastewater, agricultural run-off and urban garbage expose wild animals to resistant bacteria without any direct antibiotic use.
Foxes encounter contaminated sources while scavenging near human settlements. Waterbirds pick them up in rivers and wetlands that receive treated, and sometimes untreated, sewage.
Nine of the ten K. pneumoniae isolates shared an identical cluster of seven antibiotic resistance genes, including those conferring resistance to cephalosporins, sulphonamides, streptomycin and trimethoprim.
That the same cluster turned up across different animal species and sampling locations suggests it is moving freely through the environment.
Conter noted that although the overall prevalence of K. pneumoniae was low, at around 2%, the level of resistance within those isolates was what mattered.
“The low prevalence but high resistance levels, up to six antibiotic classes, underscores wildlife’s role as an under-monitored AMR reservoir,” he said.
What would wildlife antimicrobial resistance surveillance look like?
In practice, Conter told VaccinesWork, wildlife surveillance for AMR would “integrate passive surveillance (collecting samples from naturally deceased animals found during official wildlife mortality programmes) with standardised phenotypic and genomic testing of key indicator bacteria like Klebsiella spp. and E. coli”.
The system would involve veterinary services, environmental agencies and public health authorities working together, with annual sampling across urban and rural areas and targeted use of selective growth media for resistant strains.
The information gathered could then guide “targeted actions such as enhanced wastewater treatment to reduce antibiotic pollutant discharge, restrictions on agricultural run-off in wildlife habitats, and biosecurity measures in areas with high human-animal overlap,” said Conter.
The study was not designed to trace direct transmission between wildlife and humans, he says.
Conter says expanding the work will require longitudinal studies that connect wildlife sampling with environmental and clinical data, “leveraging high-throughput genomics for transmission tracking”, and harmonised EU-wide protocols to make the results comparable across countries.
“National One Health plans should mandate wildlife AMR surveillance in routine monitoring, with funding for integrated data platforms to inform stewardship and predict resistance hotspots,” he told VaccinesWork.